The aminoglycoside model is not hard-coded into the program. The parameters are found in the drug model database and are fully user-editable. You can tailor each drug model to fit your patient population, or you can create your own models. See the Edit drug models section of the help file for further information.
Traditional method
Initial dosing
1. Calculate dosing weight (DW)
DW = LBW + [(ABW - LBW) x CF]
where ABW = actual weight
LBW = lean body weight
CF = 40% correction factor for obesity
2. Calculate loading dose (LD)
LD = (CpTmax x Vd x kel x tinf) / (1 - e-kel x tinf)
where CpTmax = Target peak
Vd = population volume of distribution (from 3i. below)
kel = population elimination rate (from 3ii. below)
tinf = length of infusion
3. Determine initial maintenance dose (MD)
i. Calculate volume of distribution (Vd)
Vd = 0.27 x DW
ii. Calculate elimination rate (kel) from creatinine clearance
kel = 0.01 + (CrCl x 0.0024)
where CrCl = creatinine clearance
iii. Calculate ideal maintenance dose
MD = kel x Vd x CpTmax x tinf x (1 - e-kel x tau / 1 - e-kel x tinf)
where CpTmax = Target peak
Vd = population volume of distribution
kel = population elimination rate
tinf = length of infusion
iv. Calculate ideal dosing interval (tau)
tau = tinf + [-1 / kel x ln (Cptmax/Cptmin)]
where CpTmin = Target trough
CpTmax = Target peak
v. User selects practical dosage and interval
vi. Calculate expected steady-state peak & trough levels
CpSSmax = [MD / (tinf x Vd x kel)] x [(1 - e-kel x tinf) / 1-e-kel x tau)]
CpSSsmin = CpSSmax * e-kel x (tau - tinf)
Traditional method
Adjust maintenance dose using the Sawchuk and Zaske method
1. Calculate elimination rate (kel)
kel = (ln Cpk/Ctr') / time between samples
where Cpk = Measured peak level
Ctr' = Measured trough level after the dose, or extrapolated from pre-dose trough.
2. Calculate volume of distribution (Vd)
Vd = [(Dose/tinf) / kel] x {(1 - e-kel x tinf) / [Cpmax - (Cpmin x e-kel x t')]}
where Cpmax = Extrapolated peak level
Cpmin = Extrapolated trough level
tinf = infusion time
t' = hours between time Cpmin drawn and end of infusion
3. Calculate ideal dosing interval (tau)
tau = tinf + (-1/kel) x ln (CpTmax/CpTmin)
where CpTmin = Target trough
CpTmax = Target peak
4. Calculate ideal maintenance dose (IMD)
IMD = kel x Vd x CpTmax x [(1 - e -kel x tau) / (1 - e -kel x tinf)]
5. User selects practical dosage and interval
6. Calculate expected peak & trough levels
CpSSmax = [MD / (tinf x Vd kel)] x [(1 - e-kel x tinf) / (1 - e-kel x tau)]
CpSSmin = CpSSmax * e -kel x (tau - tinf)
Traditional method
Adjust maintenance dose using the 1-compartment Bayesian algorithm
1. Minimize Bayesian function
The Bayesian method uses population-derived pharmacokinetic parameters (ie., Vd and kel) as a starting point and then adjusts those parameters based on the serum level results taking into consideration the variability of the population-derived parameters and the variability of the drug assay procedure. To achieve that end, the least squares method based on the Bayesian algorithm estimates the parameters kel & Vd which minimize the following function:
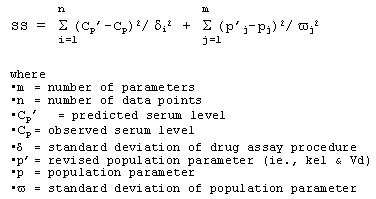
2. Calculate ideal dosing interval (tau)
Same as the Sawchuk and Zaske method
3. Calculate ideal maintenance dose (IMD)
Same as the Sawchuk and Zaske method
4. User selects practical dosage and interval
5. Calculate expected peak & trough levels
Same as the Sawchuk and Zaske method
Extended interval method
Initial dose
1. Calculate dosing weight (DW)
Same as Sawchuk and Zaske's method
2. Calculate maintenance dose (MD)
MD = DW x QDdose
where QDdose is the daily dose in mg/kg:
• Amikacin, kanamycin = 15 mg/kg
• Gentamicin, Tobramycin, Netilmicin = 5 mg/kg
3. Determine interval
Interval is based on creatinine clearance
CrCl |
Interval |
Over 60 |
24 hours |
40 - 59 |
36 hours |
30 - 39 |
48 hours |
Less than 30 |
Use traditional methods |
Extended interval method
Adjust maintenance dose
1. Determine interval
Obtain a mid-interval drug level 6 to 16 hours after the initial dose, then evaluate the interval based on the dosage adjustment nomogram.
If the 6 to 16 hour level is undetectable and the infection is not responding, consider changing to a traditional dosing method.
The three interval break points on the graphs are decay curves, produced by using a population average volume of distribution of 0.25 L/kg and an elimination rate calculated from creatinine clearances of 25, 40, and 60 ml/min for 48, 36, and 24 hour intervals respectively. The authors of the Hartford nomogram then flattened these decay curves to simplify the nomogram. The Hartford nomogram is utilized by Kinetics if your model EI dose is 7mg/kg.
|
It is important to note that the Hartford ODA nomogram is only valid for a 7mg/kg dose. An interval adjustment nomogram for the less aggressive dose of 5mg/kg/day was developed by a consensus panel. For 15mg/kg doses of amikacin multiply the drug-level scale by a factor of three. The consensus nomogram is utilized by Kinetics if your model EI dose is 5mg/kg.
|
This same consensus panel argues that the 48 hour interval should be abandoned, that patients with a CrCl < 40ml/min should be dosed by traditional pharmacokinetic methods..
Furthermore, some have questioned the validity of all ODA nomograms because they are based on one-compartment parameters derived from studies of traditional dosing methods. Some pk studies have shown that the pharmacokinetics of aminoglycosides at high doses differ significantly from those at traditional doses. Therefore, it is argued that nomograms based on an assumption of similar kinetics are invalid.
See also:

